We conducted a Double Digest Restriction Associated DNA study on Mimulus guttatus (Erythranthe guttata). The first step was collecting samples, which we did on two field trips (see Lab 3 and Lab 4 posts).
Next, we extracted DNA from samples we collected, as well as samples previously collected by Alec Chiono (see Lab 8 post).
Next, we double digested our DNA using two restriction enzymes. (See Lab 11 post). These enzymes cut up the genome into many pieces.
Next, we ligated unique DNA barcodes onto each individual (see Lab 11 post).
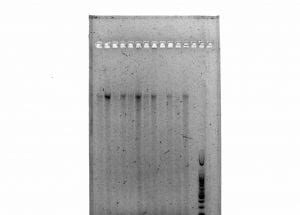
Next, we used PCR (see Lab 12) in order to 1) add a second barcode and 2) to test if our library construction was successful. Our PCR was successful, as evidenced by the results of our gel run (photo to be uploaded by Dr. Paul).
After the test PCR, we did a larger reaction that was identical and will be used for the following test. (This was the last step that we were able to do as a class.) In a perfect world, we would do the following other tests:
The next step would be size selection. Size selection selects DNA of specific sizes. Specifically, we would target approximately 400-600 bp fragments. Size selection can be done in 3 different ways, 1) PPEN prep (we have one housed in the Suni Lab at USF — #Suni), or 2) gel extraction, or finally 3) use magnetic beads to isolate the DNA fragments.
After size selection, we would then normalize our DNA samples. This means to bring all of our DNA samples to approximately the same concentration. Having equal concentrations makes more likely that equal number of DNA fragments get sequenced.
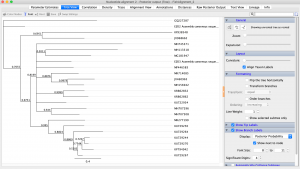
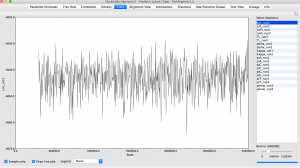
The final step is to combine all of our size selected, normalized PCR products into one vessel. We would then run these samples on any Illumina sequencer (our class would use the in-house iSeq 1000 aka Wall-e). Sequencing would take approximately 16 hours, and if successful, would generate 10s of millions of reads. These data would be run through a bioinformatics pipeline (“Help, Dr. Z!”). Ultimately, we would align these sequence data with the published Mimulus guttatus genome and call SNPs (identify SNPs in dataset).
Finally, we would use these SNPs to infer population differentiation using a metric like Fst, and assess population genetic diversity, looking at things like number of alleles, allelic diversity, etc.
Based on what I know about Mimulus guttatus, I would expect populations that are geographically divergent to be genetically divergent. Within that geographic differentiation, I would expect that populations that are ecologically divergent to be more genetically divergent.
#molecularecologyforever